Ohio University
Learning & Intelligent Systems Lab (LiSL)
Current Projects
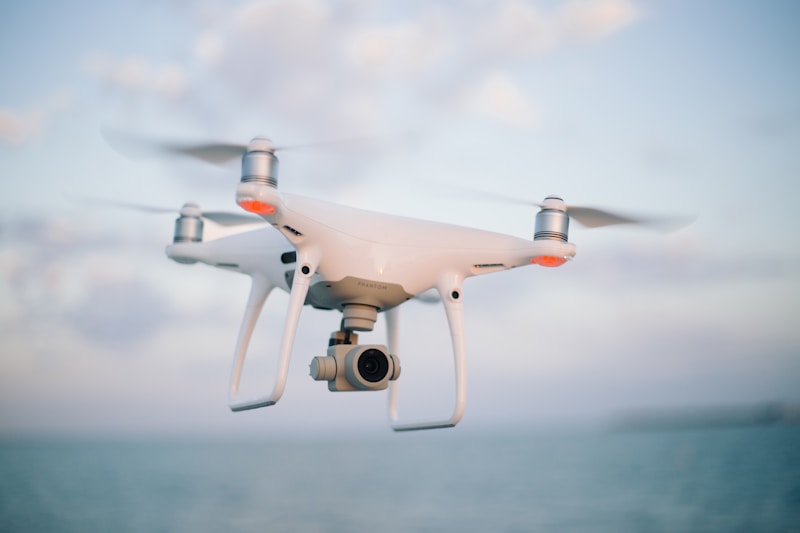
Autonomous Robots & UAVs
Methods: Deep Reinforcement Learning, Computer Vision
Sponsors: Air Force Research Laboratory (AFRL), Ohio Department of Higher Education (ODHE), NASA, Ohio University
We develop deep reinforcement learning approaches for autonomous navigation, collision avoidance, and path planning in robots and UAVs.
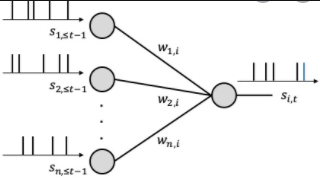
Neuromorphic Computing & Edge AI
Methods: Spiking Neural Networks, ANN-SNN Co-training, Hardware Deployment, Software-hardware Co-design
Sponsors: US Air Force, NASA, Ohio University
We develop energy-efficient spiking neural networks (SNNs) for low SWaP-C (Size, Weight, and Power-Cost) edge AI applications.
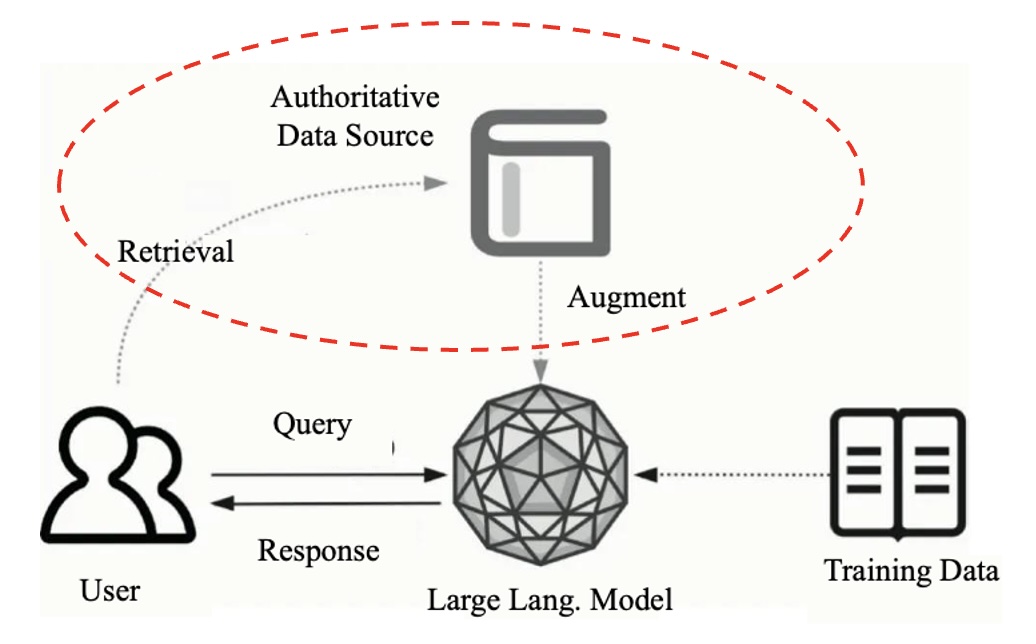
Agentic RAG and LLM Agents
Methods: Large Language Models, Agents, KV Cache, RAG
Sponsors: Flexday Solutions
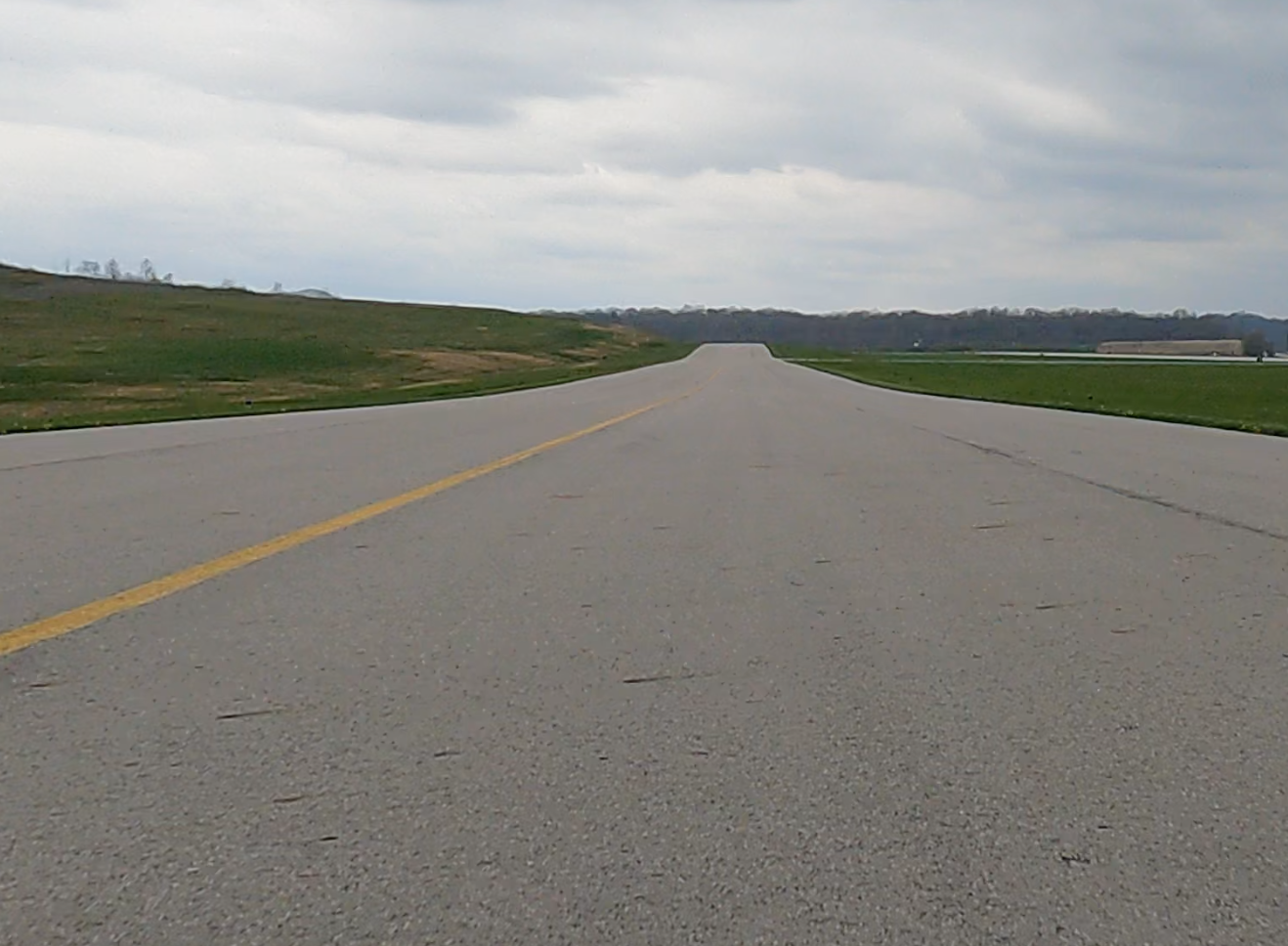
Airport Runway Monitoring (FOD Detection)
Methods: DETR, YOLO, Generative AI
Sponsors: FAA
We develop AI-based Foreign Object Debris (FOD) detection systems for airport runway safety monitoring.
Past Projects
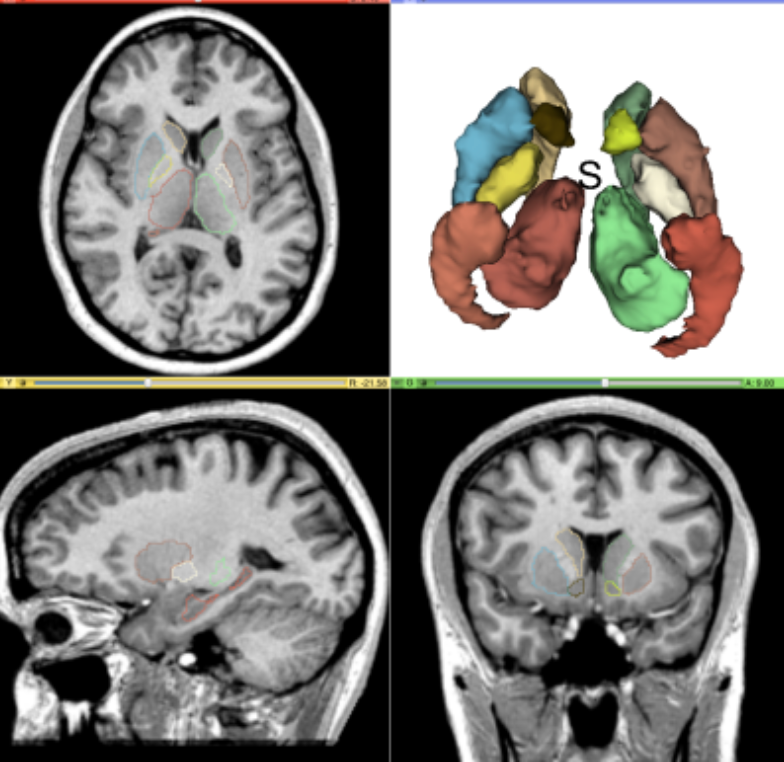
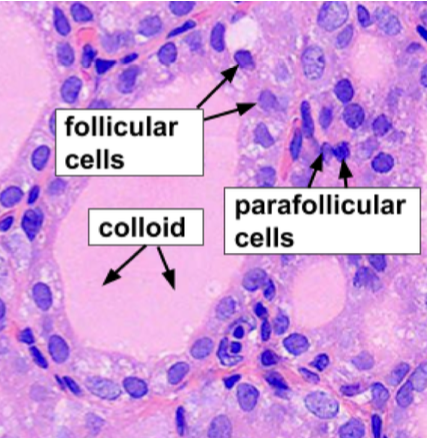
Medical Image Analysis
Methods: Deep Learning, Metric Learning, SNNs
Sponsors: University of Kentucky, NIH, Charles River Labs, Flexday Solutions
We develop deep learning and metric learning approaches for brain image segmentation and Alzheimer's disease staging.
We develop FCN-based approaches for medical image segmentation in digital pathology applications.
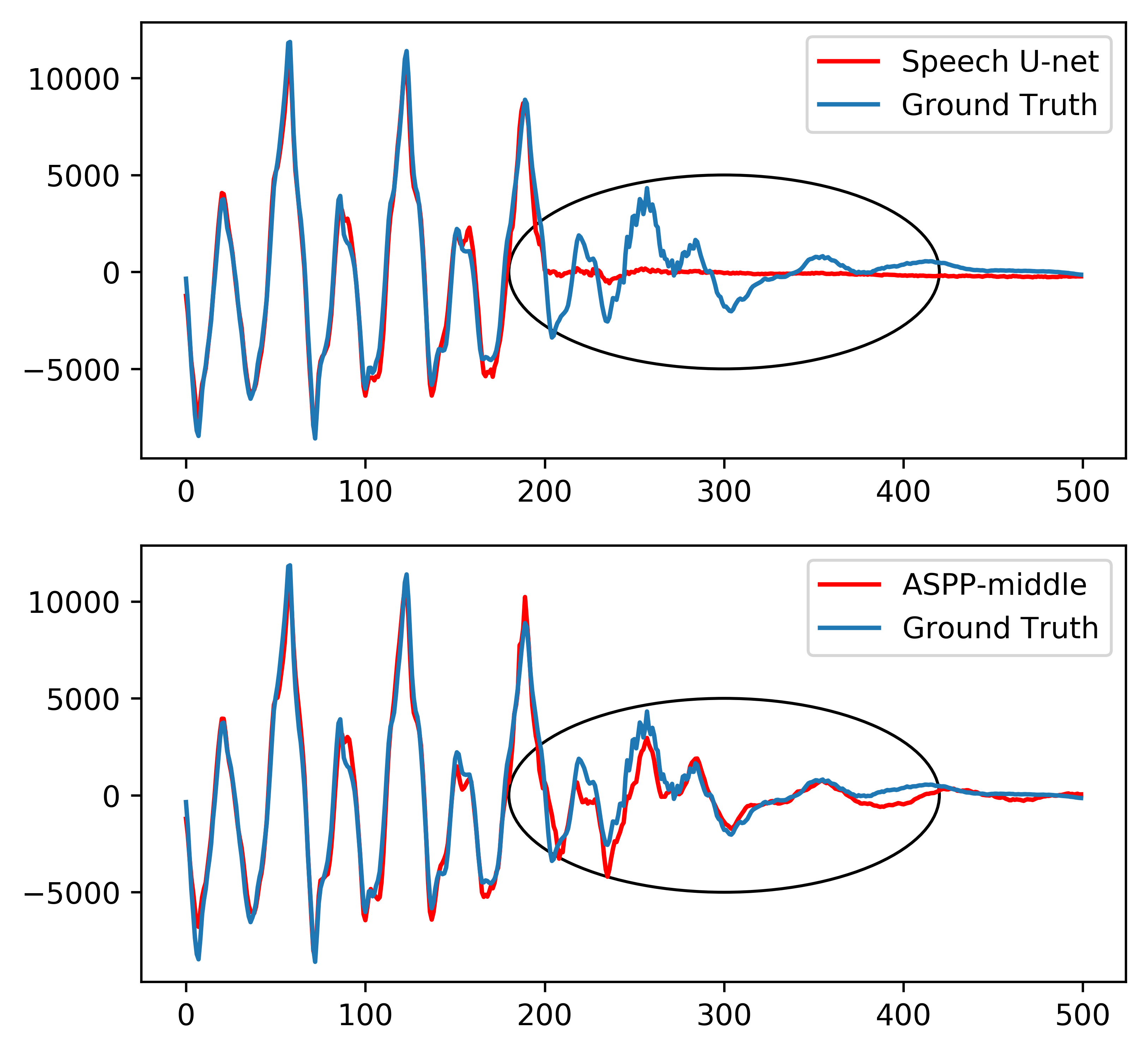
Speech Enhancement
Methods: Deep Learning, SNNs
Sponsors: Ohio University
We develop deep learning approaches for speech enhancement, speaker separation, and audio processing.
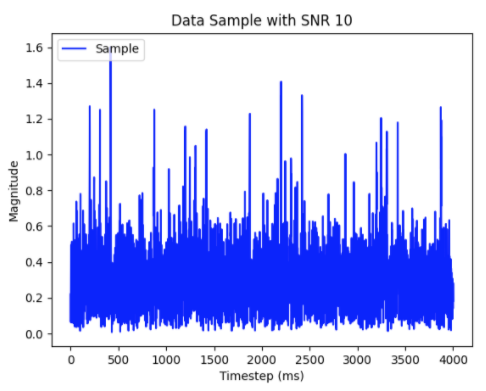
Radar Signal Processing
Methods: Network Fusion, SNNs
Sponsors: US Air Force
We develop deep learning and spiking neural network approaches for radar emitter detection and signal processing.
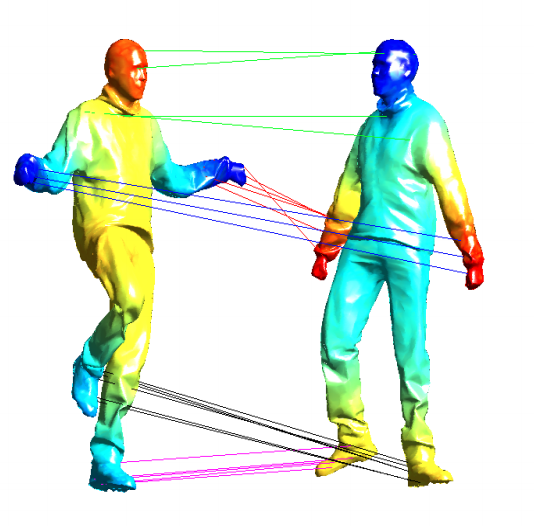
Information Retrieval
Methods: Graph Matching, Spectral Embedding
Earlier work on graph-based approaches for shape matching and information retrieval.